Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Neuroscience
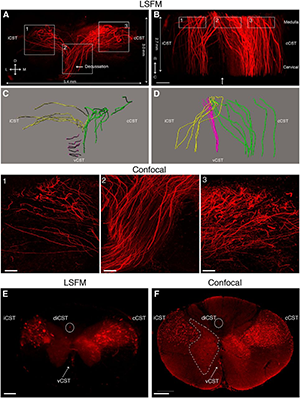
Rubrospinal tract (RST) and corticospinal tract (CST) axons can be visualized in cleared mouse spinal cord using adeno-associated viruses (AAVs). A, 3D reconstruction from serial light sheet fluorescent microscopy (LSFM) optical sections of a segment of the upper thoracic spinal cord from a mouse injected with AAV8-UbC-GFP into sensorimotor cortex to label CST axons (white). B, Dorsal view of the same 3D reconstruction as in A. C, D, Confocal microscopy, which has higher resolution than LSFM, can better distinguish individual CST axons. Boxed area in C is represented in D. E, 3D reconstruction from serial LSFM optical sections of a cleared segment of the upper thoracic spinal cord from a mouse injected with AAV2-UbC-tdTomato into the red nucleus to label RST axons (white). F, Coronal view of the same 3D reconstruction as in A. G, Projection view of coronal image stack taken with confocal microscopy from cleared spinal cord (as shown in E and F). Bracketed regions in E, F, and G refer to non-RST axons that were mislabeled during virus injection. Boxed area in G is shown at higher magnification in H. All images are taken from cleared mouse midthoracic spinal cord tissue at 4 weeks after virus injection. Scale bars: C, 80 microns; E-G, 100 microns. n = 3 biological replicates per group. iRST, Ipsilateral RST; cRST, contralateral RST. Reprinted from Soderblom et al. Eneuro 2(2).
Study | 3D imaging of axons in transparent tissue using Imaris
Drs. Jae Lee, Pantelis Tsoulfas and colleagues from the University of Miami School of Medicine developed a new technique that makes use of Imaris software and viral tracers for imaging axon trajectories in the brain or spinal cord after tissue clearing. The researchers demonstrated the new method in injured spinal cords of mouse, rat, and nonhuman primate spinal cords.
Traditionally, tracing axonal trajectory in the brain or spinal cord required laborious reconstruction of images from serial tissue sections. New tissue clearing methods combined with the right microscopy techniques now allow the tracing of axonal trajectory in intact tissue, thereby making this process much faster and, importantly, more accurate. However, most studies have used mouse models in which a subset of axons are pre-labeled at birth. By using viral tracers, the new technique gives scientists improved spatial and temporal control over their experiments.
For the study, Lee and Tsoulfas’s team used adeno-associated viruses and chemical tracers as well as tetrahydrofuran-based tissue clearing before imaging axons using light sheet and confocal microscopy. After trying many different types of software for 3D imaging analysis, they decided on Imaris.
“Imaris stood out from the rest in terms of user-friendliness and intuitive interfaces,” says Dr. Lee. “Also, the quality of the 3D videos rendered from our image stacks was significantly better than those from other programs. This video rendering capability has been very useful as we apply Imaris in our studies.”
Imaging minor axon tracts
Using Imaris, they were able to obtain a projection pattern of axons not located in the main tract of a given axonal pathway. “Such minor tracts or ectopic axons are often missed in two dimensional analyses, but the 3D videos from Imaris made these axons much more visible, allowing us to gain a much better understanding and appreciation of these axons,” said Dr. Tsoulfas. “In addition, Imaris allowed us to study the relationship between axons and other cells around the injury site in 3D, which provided a much more accurate view of the cellular organization.”
Although the researchers demonstrated the imaging approach using injured spinal cords, it can also be applied to uninjured brain and spinal cord studies. They point out that combining this approach with existing transgenic mouse lines or using it in species in which transgenic labeling is unavailable would provide researchers with enhanced flexibility in investigating axonal trajectories in multiple species with improved accuracy and speed.
Since publishing this work, the researchers have extended their experiments in the cellular transplantation area. They ultimately want to image multiple types of cells and the growth of injured axons after spinal cord injury, which will allow them to clearly distinguish between regenerating axons and collateral sprouting. They have also used Imaris in other studies to view the spinal cord vasculature in 3D.
Browse our range of related assets below...
Author: Jae Lee and Pantelis Tsoulfas, University of Miami School of Medicine
Category: Case Study
