Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Developmental Biology
Researchers led by Yu Shan Huang of National Yang-Ming University are using Imaris software to study how eyes develop in Drosophila embryos. They are particularly interested in finding out how the eye-antenna disc primordium (EADP) develops, and from where the EADP cells originate in the embryonic stage of Drosophila.
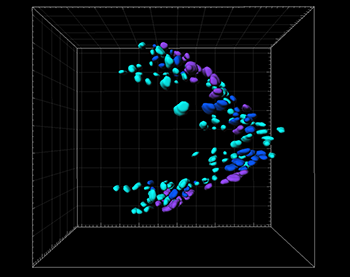
The correlation between the track duration and the spatial distribution of CD-GFP-expressing cells in the EADP. The 3D surface visualization of CD-GFP-expressing cells on the EADP from dorsal view at mid stage 16, which was the latest trackable time point. Each cell with track durations in the ranges of 0 to 63 min, 64 to 126, and 127 to 276 min, is color-coded in light blue, deep blue, and purple, respectively. Anterior of the embryo is to the left. Grid size in background is 10 µm.
Scientists want to know more about the cell dynamics of EADP formation because this would reveal important information about migration patterns, cell arrangements, and relative motions involved in this complex biological process. For more than a decade, it has been difficult to study Drosophila embryos in late embryonic stages. However, the recent development of light sheet microscopy has provided the temporal and spatial resolution necessary for long-term in vivo observation of Drosophila embryogenesis.
Visualizing large 5D data sets
The researchers used dual-illumination light sheet microscope for their study because it provides high-quality time-lapsed 3D images of live specimens. However, this technique can produce incredibly massive amounts of data. For example, a time-lapse observation lasting tens of hours might generate hundreds or thousands of gigabytes of data. To handle this massive amount of data, the researchers turned to Imaris.
“Imaris has great capability to rapidly access, visualize, and process such a massive 5D (3 spatial, one temporal, and one spectral) image data set,” said Huang. “The intuitive user interface was also helpful for efficiently quantifying and presenting our experimental results.”
The researchers genetically modified Drosophilia embryos to express both CD-GFP, a gene marker that defines EADP cells, and en-RFP, a gene marker for the posterior cells in the imaginal discs. They then imaged the embryos for 40 seconds every 3 minutes to create a 3D image stack. The resulting 5D images were processed and presented using Imaris software. The researchers used Imaris to identify and track CD-GFP and en-RFP co-expressing cells, and then created a channel to filter the co-localization signals and established the 3D isosurfaces of those cells.
Watching cells over time
“The cell tracking results provided abundant quantitative information on the positions, speeds, and signal intensities of each cell over time, so that we could compare the correlations between various factors by visualizing the target cells in different color-codes according to different parameters,” said Huang. This analysis revealed that the timing of the onset of CD-GFP expression has a certain distribution that may be related to the specific destination of EADP cells.
“Based on solid evidence from a sequence of time-lapsed 3D multi-color images with unprecedented detail, we have confirmed that EADP-engaged en-expressing cells (EECs) originate from the dorsal-lateral procephalic region in embryonic stage 13,” said Huang.
So far, their imaging analysis has uncovered the trajectories of cells that make up only a small portion of the EADP, during the embryonic stage 13 to 16. The researchers are now working to decipher the origination of other EADP cells, how the EADP folds to form a sac-like structure, and how it attaches to the brain.
Author: Yu Shan Huang of National Yang-Ming University, Taipei, Taiwan
Category: Case Study
