Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Cell Biology
Researchers led by Jordan Beach from National Heart, Lung and Blood Institute, National Institutes of Health, used super-resolution approaches together with Imaris to study the molecular mechanisms involved in non-muscle myosin 2 (NM2) assembly in migrating cells. The research revealed new information regarding how cells migrate and could provide insight into how NM2 is assembled in many other cellular processes.
NM2, the major contractile enzyme in all non-muscle cells, is closely related to the muscle motor protein that drives muscle contraction. Although both NM2 and muscle motor proteins both assemble into filaments that hydrolyze ATP and pull on actin filaments, cells can rapidly change when and where NM2 filaments form and are active. This dynamic element provides control of where and when the cell is producing contractile force and is critical for many cellular processes, such as cell migration, cell invasion, cell division, and epithelial barrier function.
Association of NM2 and actin
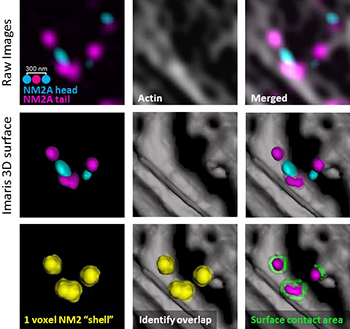
The researchers used a novel Imaris surface-surface contact XTension to determine the amount of contact between the NM2A surface and the actin surface. From the raw images (top row), they created 3D surfaces of the NM2 and the actin (middle row). The XTension created a 1-voxel “shell” around the desired surface, identified areas where the voxels in the “shell” overlap with the actin surface and then identified the percentage of the NM2 surface in contact with actin (bottom row). Courtesy of Kyle Bruun and Jordan Beach, National Heart, Lung and Blood Institute.
Beach and his colleagues were interested in studying NM2 because the cellular mechanisms governing this enzyme’s filament assembly are largely unknown. Using super-resolution imaging, they observed that the filaments repeatedly partitioned off from one another during assembly, creating multiple filaments. They hypothesized that this partitioning was driven by dynamic movements of the actin fibers to which the NM2 filaments were bound. Because the actin in this region of the cell was very dense and dynamic, quantitating the live-cell data proved difficult. For this reason, the researchers decided to conduct fixed-cell analysis using images from a structured-illumination microscopy system.
To test their hypothesis, Beach’s research team wanted to answer the question “Are NM2 filaments in the middle of the partitioning process more likely to be associated with actin fibers than randomly oriented NM2 filaments?” They note that even structured-illumination microscopy, which has a spatial resolution of about 130 nm, can only show that there is an increased likeliness that NM2 and actin are associated.
“The complexity of the particular question and the 3D nature of the analysis necessary to answer the question led us to use Imaris, which is built to answer these types of questions,” said Beach. “Working with Bitplane Application Specialist Matthew Gastinger — who was incredibly helpful — we decided the best approach was to use and update a novel surface-surface contact XTension he had been developing.”
Surface-surface contact XTension
The researchers used the Imaris surface-surface XTension to determine the amount of “contact” between the NM2A surface and the actin surface. To do this, they first used Imaris to create 3D surfaces representing the NM2 and the actin by setting a threshold for signal intensity, which determined which voxels from the original image were included in the surfaces. Applying the XTension then created a 1-voxel “shell” around the desired surface, in this case the NM2 head domain. Next, the XTension identified areas where the voxels in the “shell” overlap with the actin surface. By calculating the area of overlap divided by the total area of the shell, the XTension identified the percentage of NM2 surface in “contact” with actin.
As a control, the researchers created surfaces for regions of actin immediately adjacent to the experimental region, then dropped the NM2 surfaces randomly into those control actin surfaces. They then re-performed the surface-surface contact XTension with these non-associated areas of actin, which provided a control to show that the level of association between the NM2 and the actin in the experimental region is real and not random.
The results from the quantitative analysis performed with the surface-surface XTension supported the hypothesis that actin fiber dynamics likely play an important role in actively partitioning NM2A filaments. The study also revealed that NM2 filament assembly occurs via sequential amplification following initial nucleation and that relatively sparse nucleation events give rise to filament clusters that then populate structures present deeper in the cell and that partition frequency and filament growth rate depend on myosin light chain kinase. Overall, the study’s results provide new insights into the mechanism and spatio-temporal regulation of NM2 filament assembly in cells.
Author: Jordan Beach and Kyle Bruun, National Heart, Lung and Blood Institute, National Institutes of Health
Category: Case Study
