Cell Biology
Michael Y. Gerner, University of Washington and Ronald N. Germain, National Institute of Allergy and Infectious Diseases/NIH.
Researchers led by Michael Y. Gerner of the University of Washington and Ronald N. Germain, National Institutes of Health, developed a new tissue clearing technique that makes it possible to understand the precise location of diverse groups of cells in large tissue volumes using multiple marker combinations.
Combining this new approach with Imaris allowed the researchers to perform multiplex, high-resolution visualization of phenotypically diverse cells within organs. “To our knowledge, this is the first demonstration of quantitative phenotypic characterization of complex cell populations, such as cellular lineages and subsets, in the context of both localized microenvironments and macro-scale 3D tissue organization,” said Gerner.
This new imaging analysis approach will make it easier for scientists to study the relationships between cells and their microenvironments within whole organs. It could also benefit medical diagnostics by offering a better way to stratify patients for individualized cancer therapy, for example.
“Imaris is a robust platform for 3D and 4D visualization of large multiplex images, and using Imaris to create cell objects using Imaris for this study was a natural extension of our previous work,” said Gerner. “We have used this tool extensively in the past to analyze diverse immunological processes.”
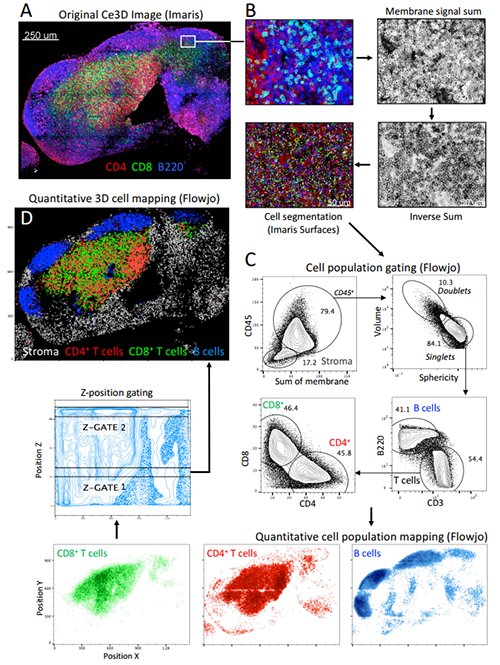
Optimizing tissue clearing
Although many approaches for tissue clearing have been developed, none are optimized for high-resolution, multiplex imaging, which requires bright fluorescence from reporter proteins while also being compatible with immunolabeling. The new easy-to-use tissue clearing technique, called Clearing enhanced 3D (Ce3D), makes tissues highly transparent while also preserving cellular morphology and protein fluorescence. It is also compatible with antibody-based immunolabeling and commonly used fluorophores.
Because Ce3D delivers high quality images and strong fluorescence signals, the researchers used the clearing method with an enhanced version of an image analysis pipeline they developed known as histo-cytometry. This imaging pipeline incorporates Imaris to accomplish 3D image segmentation and quantitative analysis of phenotypically complex immune cell subsets in 3D tissues. In the new work, the research team enhanced the histo-cytometry pipeline to perform segmentation of all the imaged cells in large 3D datasets, even cells in which the nucleus wasn’t stained.
“Imaris excels at large volume tissue visualization in three dimensions and was integral for inspecting the quality of our initial datasets for developing the clearing protocol,” said Gerner. The researchers also used Imaris-XT Channel Arithmetics to manipulate images for downstream cell segmentation and made use of the Imaris Surface Creation module for 3D segmentation of individual cells and to export statistical data for further statistical analysis.
Bridging the gap between flow cytometry and imaging approaches
Ce3D combined with histo-cytometry generates a quantitative profile of cells and tissues that can be used for additional statistical analysis. This capability makes this the approach useful for studying the locations of specific cell populations and where they perform critical functions, as well as examining the mechanisms that promote these spatial processes.
“By providing a comprehensive strategy for whole-organ cellular and molecular level quantitative imaging and analysis, our technology bridges the disconnect between dissociation-based flow cytometry and imaging-based methods, and is thus of significant value to diverse fields of basic science, translational medicine and clinical diagnostics,” said Gerner.
Using Imaris and computational imaging, the researchers have made new discoveries about the distribution and mode of function of multiple immune cell subsets within lymphoid and non-lymphoid tissues. This includes the spatial distribution and function of antigen-presenting cells during homeostasis, immunization, and infection; localized immunosuppressive activity of T regulatory cells during autoimmune activation; and the spatial positioning and function of CD8+ T cells during chronic viral infection and HIV.
The researchers are continuing to improve their imaging analysis methods and apply these approaches in new studies. They are now using Imaris to study the mechanisms that drive adaptive immune response heterogeneity and to visualize immune cell positioning and functionality in tumor tissues and inflamed peripheral tissue sites during infection.
Research Paper: Li W, Germain RN, Gerner MY. 2017. Multiplex, quantitative cellular analysis in large tissue volumes with clearing-enhanced 3D microscopy. Proc Natl Acad Sci U S A. 114(35):E7321-E7330.


