Tissue Clearing
Hu Zhao and colleagues at Texas A&M have developed a new tissue clearing method that renders nearly all tissue types transparent while preserving endogenous fluorescence. By combining the new method with Imaris image analysis software, they were able to clear the tissue of a whole mouse and reconstruct its intact jawbone, teeth, femur, and knee joint in 3-D.
Improved tissue clearing
Tissue clearing makes opaque tissues transparent, allowing various organs and body parts to be visualized in 3D. However, individual tissue clearing protocols typically only work for certain kinds of tissues. Hard tissues such as bones and teeth are particularly difficult to clear, especially if endogenous fluorescence needs to be preserved.
The researchers developed a new tissue clearing method, called polyethylene glycol (PEG)-associated solvent system (PEGASOS), that works for all types of tissue, including bones and teeth. It can be combined with various imaging methods, such as confocal microscopy, two photon microscopy and light-sheet microscopy to create 3D reconstructions.
“We used Imaris to do the 3D visualization and analysis for almost all the whole-body images,” said Zhao. “Imaris provides modules that are user-friendly and easy to operate, including 3-D rendering, video making, image segmentation, and neuron tracing. The processing speed is also faster due to optimized hardware support.”
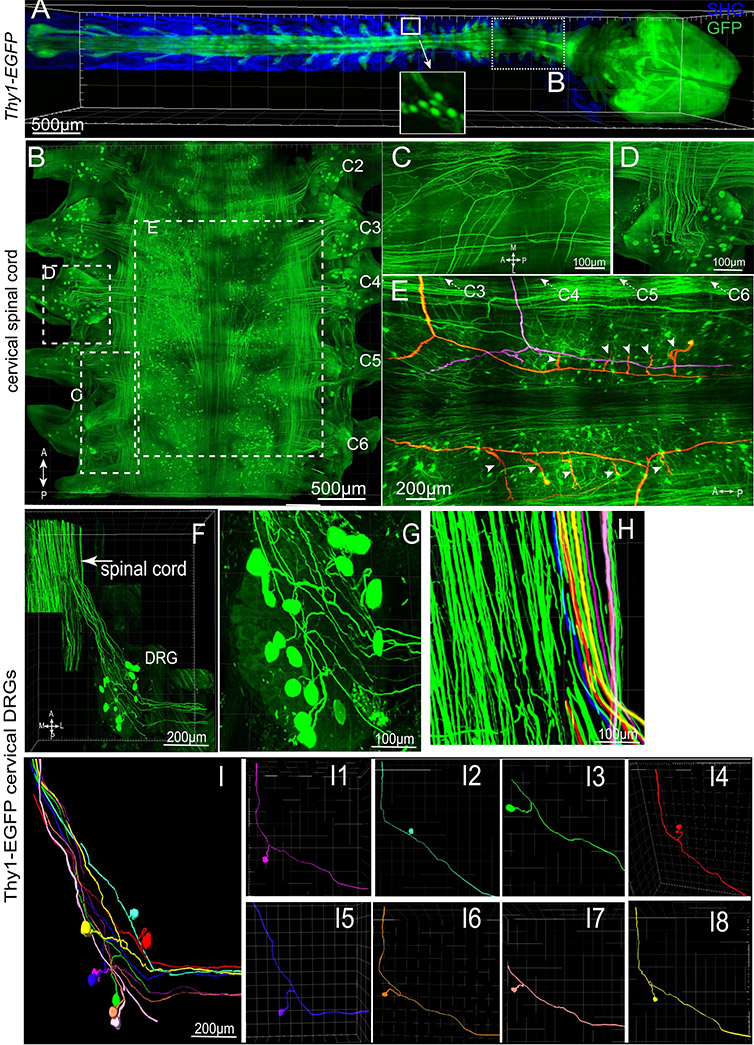
Figure 1. PEGASOS enables visualization of connections between CNS and PNS by imaging through the vertebrae. An adult Thy1-EGFP mouse (2 months old) was cleared following PEGASOS recirculation procedure and the vertebrae and brain were dissected for imaging. (a). The intact CNS together with DRGs was imaged with a 5X/0.16 objective. Boxed area is enlarged in the inlet to show individual DRG neurons. (b) The cervical vertebrae (dotted box in a) was re-imaged with a 10X/0.45 objective to visualize cervical DRGs C2-C6. Optical sections were acquired at boxed regions to show the multiple central axons (c) and DRG neurons (d). (e). Tracing of individual central axons. Each central axon (highlighted with color) gives rise to two daughter branches and then to multiple collateral branches (arrowheads). The entire tracing length is over 2 cm. (f-i). The C4 DRG was re-imaged with a 20X/0.95 objective to reveal connections between DRG neurons and the spinal cord. (g). Enlarged view shows neurons within the C4 DRG. (i). Eight pseudo-unipolar neurons were individually identified and artificially labeled with different colors (i1-i8). Their central axons can also be individually identified within the spinal cord (h). A, anterior; P, posterior; L, lateral; M, medial. Reprinted from Jing D et al, Cell Res. 28(8):803-818 under Creative Commons Attribution 4.0 International License.
When the researchers used their PEGASOS method to clear the whole body of adult mice, they observed high transparency for body parts including the entire head and brain, thorax, and both front and hind legs. Complete transparency was also seen for internal organs including the brain, liver, kidney, and spleen. In addition to creating multiple 3D renderings of various body parts and organs, the researchers were able to image the intact mouse brain at sub-cellular resolution, trace individual neurons and axons over a long distance, and visualize dorsal root ganglions directly through vertebrae.
Viewing cleared samples in 3D
“We found the 3D rendering in Imaris, especially mapped color display, was a good way to restore the structure of our samples — including neurons, axons, or blood vessels — within an organ,” said Zhao. “The Imaris animations helped display the 3D structure more intuitively while Surface rendering was useful for outlining areas of interest.”
They also used image segmentation to view the surface of areas of interest and the statistics function to count the numbers of the cells in 3D images.
“Our results suggest that the PEGASOS method provides a useful tool for general biomedical research in not only neuroscience but also various kinds of research fields,” said Zhao.
The researchers are working to improve the PEGASOS method to reduce high autofluorescence in some specific tissues and tissue shrinkage caused by the fully dehydration process. “We will use Imaris for our future work for 3D display, 3D quantification, and analysis for our image stacks,” said Zhao. “We believe there are still many other outstanding Imaris functions that could be used, and we will try them in our future work.”
The researchers used Imaris software to image the vasculature of a whole mouse brain that had been cleared using the PEGASOS method. The images were acquired using a tiling light sheet microscope. Courtesy of Hu Zhao, Texas A&M.
Research Paper: Jing D, Zhang S, Luo W, Gao X, Men Y, Ma C, Liu X, Yi Y, Bugde A, Zhou BO, Zhao Z, Yuan Q, Feng JQ, Gao L, Ge WP, Zhao H. 2018. Tissue clearing of both hard and soft tissue organs with the PEGASOS method. Cell Res. 28(8):803-818.


