Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Cell Division
Valérie Lobjois and her colleagues at ITAV are using Imaris to gain insight into cell division dynamics in 3D models of microtumors. Deciphering the mechanisms involved in regulating the cell cycle and cell proliferation is fundamental to understanding how misregulation of these processes contributes to tumors growth. In fact, the study of these mechanisms has already contributed to improved targeted therapies for cancer.
Since cell cycle control mechanisms depend on cell-cell and cell-extracellular matrix interactions, the researchers used 3D tumor models called multicellular tumor spheroids that integrate these interactions. Similar to the microregions found in a tumor, the multicellular tumor spheroids display gradients of proliferating cells in the outer cell-layers whereas the inactive quiescent cells are located more centrally.
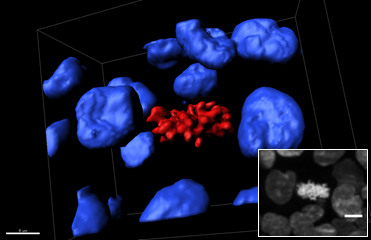
This 3D reconstruction shows a region of a SPIM z-stack of a mitotic cell located on the surface of a spheroid expression H2B-hcRed. The blue isosurfaces are interphase nuclei, and red isosurfaces indicate mitotic condensed chromosomes (scale bar: 7 microns). The corresponding SPIM frame is shown in the inset (scale bar: 15 microns).
Although studying ongoing mitosis in a 3D integrated cellular context is of great interest, scientists have only a basic understanding of the spatiotemporal dynamics of tumor cell proliferation in complex 3D systems such as spheroids. One reason for this knowledge gap is that studying the 3D dynamics of living cells in spheroids larger than 300 microns in diameter is technically challenging.
Cutting down on phototoxicity
Methods such as laser scanning microscopy are too phototoxic to use on such large samples, so the researchers decided to try selective plane illumination microscopy (SPIM). This microscopy technique uses a sheet of light to illuminate the sample, with high-resolution images recorded one plane at a time.
Using a homemade SPIM, the researchers imaged live spheroids expressing histone H2B fluorescent nuclear reporter protein and then made 3D reconstructions with Imaris. Extracting isosurfaces of the fluorescence intensity allowed visualization of individual fluorescent objects in a z-stack encompassing a mitotic cell.
“We found the Imaris option to conduct semi-automated or fully manual 3D reconstruction particularly useful, as was quantifying morphology parameters using isosurfaces,” said Lobjois. “We also liked the automatic scale bar and the ease with which we could make movies.”
Imaging deep inside spheroids
During mitosis, the cell’s microtubule network forms a mitotic spindle that aligns the duplicated and condensed chromosomes in the cell’s center and then moves a set of chromosomes to each pole, around which the daughter cell forms. The researchers could visualize the nuclei in nearly the whole volume of live spheroid by rotating the sample in the SPIM. In these 3D reconstructions, the mitotic cells with condensed chromosomes were easily and clearly identified within the spheroids.
“We demonstrated that SPIM allows live cell division imaging deep inside spheroids,” said Lobjois. “Our data showed that 3D dynamic imaging of spheroids by SPIM could be applied to evaluate the activity of putative anti-tumor drugs.”
Since the publication of this study, the researchers have used SPIM and Imaris to quantify cell division, the duration of the different phases of cell division, and cell division orientation with respect to the spheroid surface, in 3D reconstructions of metaphasic cells created using time-lapse SPIM images.
