Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Light Sheet Microscopy
Ryann M. Fame, Hazel Sive, and colleagues, Whitehead Institute for Biomedical Research and Massachusetts Institute of Technology
Ryann M. Fame, Ph.D., and her colleagues in the laboratory of Hazel Sive, Ph.D., at the Whitehead Institute for Biomedical Research and Massachusetts Institute of Technology are studying the dynamics of cerebrospinal fluid (CSF) in zebrafish embryos. Using Imaris, they were able to image CSF flow in three dimensions, revealing new details about this fluid’s movement and providing clues about its possible role in early brain development.
The brain contains a communicating network of cavities called ventricles that are filled with CSF. Scientists think that CSF likely plays an important role in brain development, since it interacts with neural stem cells during development and contains growth factors, neurotransmitters, and other signaling molecules. Some investigators hypothesize that the dynamic properties of CSF movement may restrain locally produced factors to specific regions of the developing brain. The flow of CSF might also deflect primary cilia lining the ventricles and convey cell polarity information to the developing brain
“Until now there has been no study of in vivo CSF dynamics between ventricles in the embryonic brain,” said Fame, a postdoctoral associate. “Our study provides critical information to underlie future studies of CSF component distribution and fluid properties.”
Seeing flow in 3D
The researchers studied CSF velocity between ventricles during two stages of zebrafish development. Although similar experiments have been conducted in zebrafish, CSF flow hasn’t been imaged in three dimensions previously. “Because CSF flow goes in all dimensions, we determined that a full volumetric analysis would be more appropriate,” said Fame. “Therefore, we used the 'surface creation' and 'volume/voxel calculation over time' functions of Imaris for this study.”
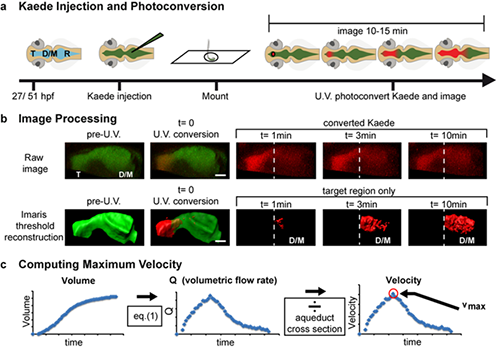
Experimental workflow. a) Schematic representation of rhombencephalic ventricle Kaede injection and confocal photoconversion and imaging of zebrafish ventricles. b) Workflow for image processing. Raw images were collected and then thresholded in Imaris software so that red (photoconverted) Kaede was only detected after photoconversion and was not detected in the unconverted target ventricle immediately after photoconversion. Then the volume of the unconverted target ventricle was calculated over time by Imaris software. Dashed line indicates telencephalic-to-mesencephalic/diencephalic boundary. c) Workflow for data analysis. From volume measurements of the target ventricle, volumetric flow rate (Q) was calculated. Q was smoothed using a 9-point moving-window average and then converted to linear velocity by dividing by the cross-sectional area of the smallest part of the aqueduct between the two ventricles. These CSF dynamics were reported as the maximum velocity between the two ventricles (vmax). T telencephalic ventricle, D/M diencephalic/mesencephalic ventricle, R rhombencephalic ventricle, hpf hours post-fertilization, V volume, Q volumetric flow rate, v velocity. Scale bar: 50 microns. Reprinted from Fame RM et al, Fluids Barriers CNS 13(1):11.
The researchers also used selective plane illumination microscopy (SPIM) to acquire full volumetric images of the zebrafish brain ventricular system, including the spinal canal. This fluorescence microscopy technique, which uses a focused light-sheet to illuminate the specimen from the side, was critical because the zebrafish yolk scatters light and obscures small details. Using Imaris, the investigators further analyzed the images to obtain static volumetric measurements of the ventricular system.
The researchers also used SPIM for high-speed imaging of central nervous system cilia. SPIM provided the high-speed, deep fluorescence imaging capabilities necessary to image the cilia movement.
The image analysis revealed directional movement for embryonic CSF in the zebrafish model during the time when most neurons are developing. High-speed imaging showed that brain cilia are not motile during the early larval stage and are thus probably not involved in directing CSF movement. The researchers also found that directional movement is partially dependent on heartbeat. Overall, the findings suggest that CSF components may be compartmentalized and could contribute to specialization of the early brain. CSF movement may also provide directional mechanical signaling.
