Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Immunology
Natural killer cells are a critical part of the innate immune system and are important for controlling infection and tumor growth. To respond to infection, these cells must be fully activated through an interaction with other immune cells. Researchers led by Dr. Janine Coombes, who was a postdoc in Dr. Ellen Robey's lab at the University of California-Berkeley, recently used Imaris to study these interactions and their regulation.
The interactions necessary for natural killer cell activation are regulated by the natural killer cells ability to navigate through, and correctly localize, within complex tissue environments. Dr. Coombes, who is now at the University of Liverpool in the United Kingdom, explains that understanding how natural killer cell motility and function are regulated in complex tissue environments could allow scientists to fine tune natural killer cell activities in a variety of disease settings.
The researchers used Imaris to analyze the relationship between the speed at which natural killer cells migrated, and their proximity to structural features in the lymph node. “Imaris allowed for easy and intuitive visualization of complex 4D data sets, the flexibility to perform automated or manual analysis of cell motility parameters, and great data presentation tools,” Dr. Coombes says.
The researchers acquired two-photon images of lymph nodes from mice, scanning the tissue every 30 seconds for 10 to 30 minutes, and obtaining image volumes that were typically 172 x 143 x 30 to 200 microns. They then used the tracking algorithmsfunction in Imaris to determine the position of each natural killer cell over time. After which, they defined the location of collagen at each time point by using Imaris to create an isosurface and filling the entire collagen isosurface with 1-micron diameter spots. For each cell At at each time point, they calculated the distance from each natural killer cell to the nearest collagen spot. They also determined the interval speed based on the position of the same natural killer cell in successive time points.
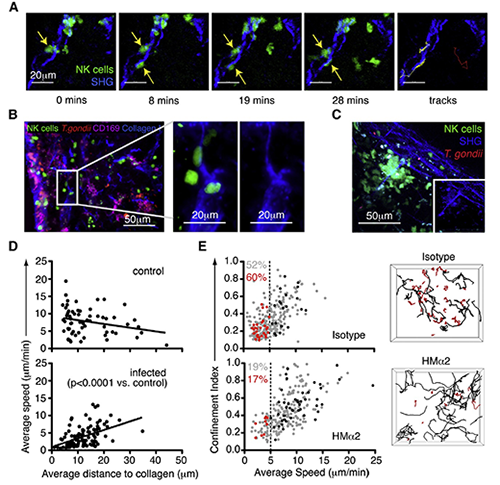
(A) Individual time points and tracks from a TPLSM movie showing two natural killer cells (green, yellow tracks) interacting with a collagen fiber (second harmonic [SHG], blue) in the cervical lymph node 24 hours after earflap infection. The track of a natural killer cell that does not interact with collagen is shown for comparison (red). (B) Analysis of cervical lymph nodes 24 hours after earflap infection by epifluorescence microscopy. CD169 staining is shown in purple, collagen type I staining in blue, natural killer cells in green, and T. gondii in red. The panels on the right are close-up views of the boxed region. (C) Still image from a TPLSM movie showing a stable swarm of natural killer cells (green) centered around a collagen fiber (second harmonic, blue). Inset image is from the same movie but with natural killer cell signal removed. (D) TPLSM analysis comparing the average speed of natural killer cells to their average distance to the nearest collagen (second harmonic) structure over the course of an imaging run is shown. Each data point represents an individual natural killer cell. (E) TPLSM analysis of natural killer cell motility in cervical lymph nodes from Ncr1GFP/+ mice infected in the earflap with T. gondii 24 hours previously, and administered anti-CD49b (clone HMa2) or an isotype control 4 hours previously. Graphs show the average speed and confinement index of individual natural killer cells. Data from an individual run are highlighted in red (cells migrating at <5 mm/min) and black (cells migrating at > 5 mm/min). These correspond to the tracks of individual natural killer cells shown in the right-hand panels. Numbers indicate the percentage of natural killer cells migrating at <5 mm/min (red for highlighted imaging run, and gray for all depicted imaging runs). Image reprinted from Cell Reports, Volume 2, Issue 1, 124-135). Image and video courtesy of Dr. Janine Coombes, University of California-Berkeley.
“We were able to show that proximity to collagen only affected natural killer cell motility in inflamed lymph nodes, lending support to our hypothesis that collagen acts as a danger signal to natural killer cells, retaining them in inflamed tissues where they are required to fight infection,” Dr. Coombes says.
The researchers continue to use Imaris to investigate mechanisms by which pathogens can alter immune cell motility in a variety of different tissue environments, contributing to the spread of infection. For example, they have also used the software to describe the movement of Toxoplasma gondii infected neutrophils in the small intestine (Coombes et al. 2013).
Research Paper: Coombes JL, Han SJ, van Rooijen N, Raulet DH, Robey EA. 2012. Infection-induced regulation of natural killer cells by macrophages and collagen at the lymph node subcapsular sinus. Cell Rep. 2012 Jul 26;2(1):124-135. .
Author: Dr. Janine Coombes and colleagues, University of California-Berkeley
Category: Case Study
