Virology
Researchers led by Dr. Seema Lakdawala from the NIH National Institute of Allergy and Infectious Diseases, USA, used Imaris software to observe influenza A replication dynamics in three dimensions. Their findings provide information that could lead to new targets for antiviral therapies.
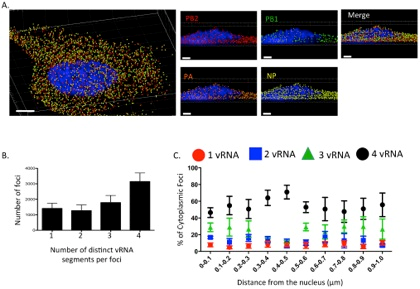
This 3D rendering generated using Imaris software shows a cell eight hours post infection with an influenza virus with four influenza viral RNA segments identified: PB2 Quasar 570 (red spots), PB1 Alexa-Fluor 488 (green spots), PA Cal Fluor Red 590 (orange spots), and NP Quasar 670 (yellow spots) (A). The DAPI-stained nucleus is labeled in blue. An enlarged region of the 3D image was tilted 90° in the z-direction to provide an axial-view of the cytoplasmic foci and nucleus. Scale bars are 4 microns in the whole cell image and 2 microns in the rotated images. The total number of foci containing 1, 2, 3, or 4 viral RNA segments was quantified within a given cell (B). Each bar represents the average from 3 independently analyzed cells with standard error indicated. The distance from the nucleus for each focus from four independent cells was calculated (C). The proportion of foci containing 4, 3, 2, or 1 distinct vRNA segment with a given range from the nucleus is represented graphically as a scatter plot. Each spot is an average from 4 independent cells and the standard error is indicated. Reprinted from Lakdawala et al., PLoS Pathog 10(3): e1003971.
Influenza A viruses cause seasonal epidemics and contain eight single stranded RNA gene segments. Influenza A can sometimes transform into strains with pandemic potential, such as H1N1 in 2009 and H7N9 in 2013, when a subtype that typically infects humans and one that typically infects animals co-infect the same cell, allowing viral RNA from the two strains to be mixed through reassortment.
Replication of the viral genome occurs in the host cell’s nucleus, and progeny viral RNA segments must travel through the cytoplasm to the plasma membrane before exiting the cell via budding. The dynamics of intracellular viral RNA assembly into progeny viral RNA segments isn’t understood, so Dr. Lakdawala and her research team set out to visualize the process.
The when, where, and how of viral RNA assembly
The researchers first studied RNA assembly using fixed cells, finding evidence that influenza viral RNA segments assemble into a single copy containing all eight segments during transport in the cytoplasm. To determine where, when, and how assembly occurs, they used Imaris and a fluorescently labeled virus to visualize viral RNA transport during a productive infection.
The researchers specifically wanted to analyze viral RNA assembly in the cytoplasm of single cells. To do this, they used the Imaris contour function to define individual cells within the field of view, masking the nucleus to ensure analysis of influenza viral RNA segments only in the cytoplasm. They then used Imaris colocalization analysis to assess the Pearson colocalization coefficient between pairs of influenza viral RNA segments. The Pearson colocalization coefficient quantifies the degree of colocalization between fluorophores.
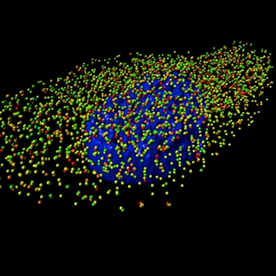
A 3D rendering generated using Imaris software of an MDCK cell infected with influenza A/WSN/1933 (H1N1) virus with four influenza viral RNA segments identified by fluorescent in situ hybridization: PB2 (red spots), PB1 (green spots), PA (orange spots), and NP (yellow spots). The nucleus is labeled in blue based on DAPI staining. Reprinted from PLoS Pathog 10(3).
The investigators also used the Imaris Spots segmentation and manual tracking to create Spots objects for each segment of viral RNA in the cytoplasm and then merged them to determine the RNA composition of each Spots object of cytoplasmic focus. “We observed that not all cytoplasmic foci contained all four viral RNA segments we stained for, demonstrating that influenza viral RNA segments are not exported from the nucleus as pre-formed complexes of all eight viral RNA segments,” explains Dr. Lakdawala.
They then used the Imaris Distance transformation XTension to determine the composition of cytoplasmic foci near the external nuclear periphery, as defined by the DAPI signal, and found that influenza viral RNA segments were not exported as individual segments, as previously suggested.
“Our data suggest that viral RNA segments are exported from the nucleus as subcomplexes that undergo additional assembly en route to the plasma membrane through dynamic fusion events of viral RNA-containing cytoplasmic foci,” says Dr. Lakdawala. “Our observations have broad implications for understanding the intracellular requirements behind reassortment of influenza viruses.”
As a next step, the researchers want to determine if there is a preference for certain segments of influenza viral RNA to form subcomplexes that undergo further assembly in the cytoplasm. For this, the researchers are determining the exact geographical location of RNA segments within the cell using the data from Imaris on the composition of each cytoplasmic focus, distance to the nucleus, and the distance to the contour made for each cell.
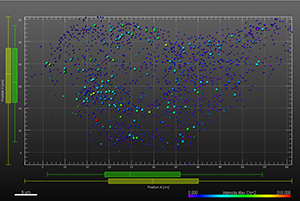
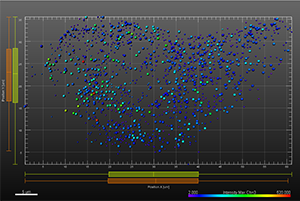
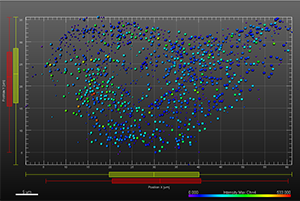
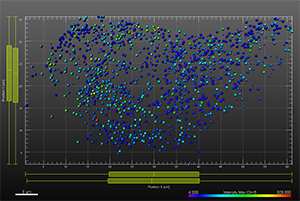
The additional images were created by the Bitplane team based on the original image data curtesy of Dr Lakdawala. Imaris Spots segmentation and statistical colour coding was used for visualization of influenza A viral RNP complex. Imaris 2D Vantage Plots illustrate the spatial distribution between encode the viral RNP proteins segmented as Spots objects: PB1 (green), PA(orange), PB2 (red), and NP (yellow). For better visualization, the colour of the objects indicates the Intensity maximum of the corresponding channel.
Research Paper: Lakdawala SS, Wu Y, Wawrzusin P, Kabat J, Broadbent AJ, et al. 2014. Influenza A Virus Assembly Intermediates Fuse in the Cytoplasm. PLoS Pathog 10(3): e1003971.







