Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
Welcome to the Imaris 9.6 release notes. Please have a look to the overview of Imaris Release notes for information about prior released features and fixed bugs.
Version Date: July 21, 2020
The Imaris 9.6 focuses on classification or labeling of objects into “classes” to distinguish these classes for downstream analysis. For example one might classify spots inside cells into ClassA and spots outside cells into ClassB to distinguish the two classes.
To facilitate object classification and analysis of classified objects we have introduced several new features in Imaris 9.6.
Another major change in Imaris 9.6 is much improved mouse behaviour in the 3DView. Navigation and Selection have been combined into a single mode removing the painful need to press Esc for the mode switch that users had to learn to do in previous Imaris versions.
A third major change that you will notice immediately is the new look of the camera toolbar in 3DView.
In Imaris 9.6 within the creation wizards for Spots and Surfaces it is possible to set up classification rules based on the statistics values that Imaris computes for each object.
The top part of the Classification User Interface lets you set up the number of classifiers to be applied, the “input” for each and the type. The “Input” drop-down let’s you choose which objects to classify when multiple possibilities are available. For example when you have a time series with spot tracks you can choose to classify tracks based on their track statistics or individual spots based on their spot statistics. Furthermore when you add a second row to add a second classifier you can choose as input one of the classes that were set up in the first row.
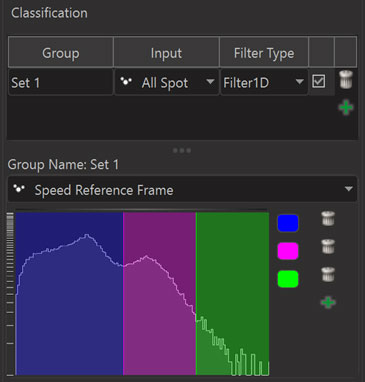
Using the Filter1D one can split objects into classes by specifying thresholds for a specific single statistics value. Using this type of classifier one can for example easily assign the objects with a low “Speed Reference Frame” value to one class and objects with larger “Speed Reference Frame” values to other classes.
Using the Filter2D it is possible to split objects into classes based on two statistics values as shown in the following screenshot.
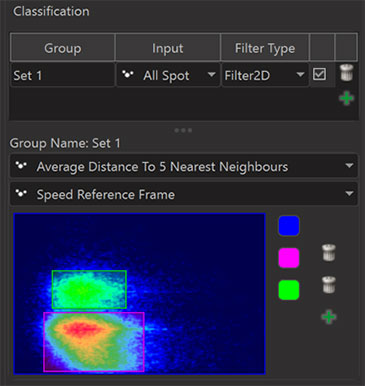
In Imaris 9.6 classes in the Filter2D dialog can be defined using rectangles only. If you encounter situations where other shapes would be preferable please let us know. We built the Filter2D dialog as a first step in a direction that we can continue in future Imaris releases.
The third possibility for automated classification is to train a Machine Learning Classifier (ML). The dialog for machine learning looks as follows before training.
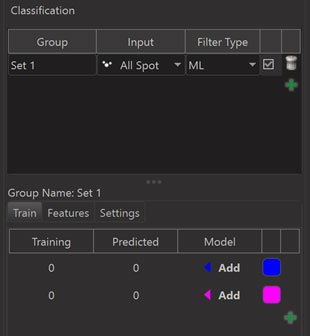
You can select objects in the 3D View and then click on an “Add” button for the class that the selected objects should be assigned to. This adds them to the “Training Set”.
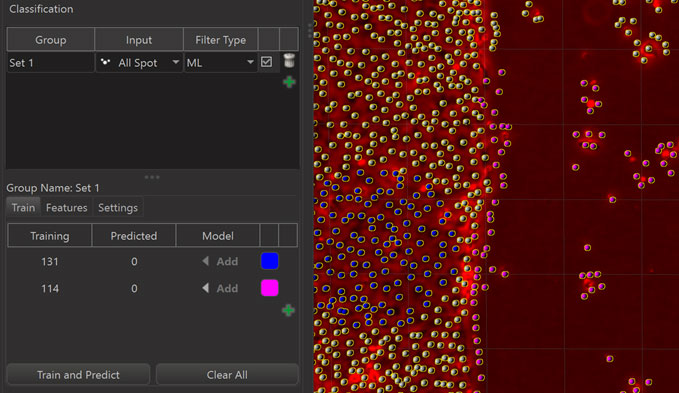
Once some useful example objects have been added to the Training Set the actual training and prediction of the machine learning algorithm happens when the “Train and Predict” button is pressed. As a result of this the class of each object that was not in the Training Set is predicted by the algorithm.
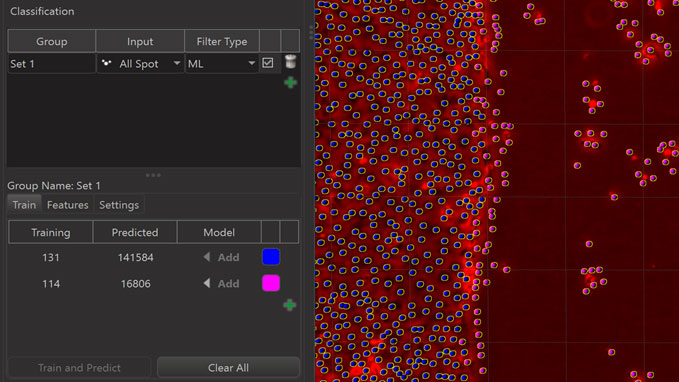
A very efficient approach to train the machine learning classifier is to start with a relatively small training set that can be created quickly and see what the algorithm can predict from that. This first prediction will likely misclassify some objects. If you then correct the classification of those misclassified objects by selecting them in the 3D view and adding them to the class you want them to be in you provide valuable information to the machine learning algorithm and the algorithm can “learn efficiently” when you press “Train and Predict” again. Repeating this cycle of prediction, correction, and training can get you toward the desired result efficiently.
Automated object classification in Imaris takes the statistics values that Imaris computes for each object as input to compute classes based on those statistics values. Imaris computes a large number of statistics values for each object: intensity statistics of each channel within the object, volume and shape of the object, speed of tracked objects and shortest distances to other objects. In Imaris 9.6 we have added some more statistics values that can be useful for the purpose of object classification. The new values are the following:
The SpotsAverageDistanceToNearestNeighbours statistics provides useful density measurements for the spots. Where the average distance is large, density is low. These statistics can be useful for discriminating regions with different spot densities.
For the machine learning classifier to have a good feature set Imaris computes a large number of features for machine learning specifically. These features do not appear in the normal set of statistics values. For most purposes it should not be necessary to look at the list of statistics values computed for machine learning. The list of machine learning statistics is visible only on the Settings tab of the machine learning user interface.The list is available mainly to allow experts to look “under the hood” and see which values the machine learning classifier gets as input.
For the interested reader we recommend the following paper that describes a machine learning approach similar to the approach implemented in Imaris: Ranzato, M., et al. "Automatic recognition of biological particles in microscopic images." Pattern recognition letters 28.1 (2007): 31-39. Ranzato et al. make use of multi-scale features invariant to shift and rotation as described originally by Koenderink and van Doorn, "Representation of local geometry in the visual system." Biological cybernetics 55.6 (1987): 367-375.
For classification of Spots Imaris computes Gaussian derivatives up to second order at a range of scales from 1 times the spot radius up to 8 times the spot radius. This creates a large set of shift and rotation invariant features that describe the local image intensity around a spot up to second order. In the list of machine learning statistics the names of these features all start with “JET”. You can think of these features as describing the local “intensity landscape” of each channel at the position of the spot in terms of how high, how steep and how curved the landscape is. In addition to the Gaussian derivatives of the original channels Imaris also computes features that describe the local “texture” of an image around a spot. To do so it computes a binary image capturing the positions of local extrema in the original channel and from this binary image it computes the same Gaussian derivatives as for the original channel intensities. In the list of machine learning statistics the names of these features all start with “TEX”. Both sets of features together provide a lot of useful features for a machine learning classifier.
For machine learning classification of Surfaces Imaris uses the existing statistics values and in addition computes more values that provide additional intensity and shape information. The intensity of each channel on the boundary of the surface is computed for machine learning. To provide additional shape descriptors Imaris computes the position and radius of the biggest sphere that fits entirely within the surface and derives a number of features from this. The simplest is the radius of the biggest sphere itself. Imaris also computes “relative weights” of the surface in shells around the center of the biggest sphere as well as the “relative weights” of the surface in shells around the center of mass. Two different types of “weights” are computed. One uses the volume inside the surface as “weight”. The other uses surface area as “weight”. In the list of machine learning statistics the names of these features are ShapeVolumeHistogramOnCenter ShapeAreaHistogramOnCenter, ShapeVolumeHistogramOnBFS, and ShapeAreaHistogramOnBFS. These values are by construction shift, rotation, and scaling invariant. The original Surfaces statistics together with the machine learning statistics provide a lot of information about the shape of a surface and can be very useful for a machine learning classifier for surfaces.
In Imaris 9.6 the Spots, Surfaces, Filaments, and Cells components of Imaris all have a tab “Edit Labels” that lets users manually label objects. This user interface replaces the labeling user interface that was previously on the camera toolbar of the 3D View.
The top of the “Edit Labels” tab is used to set up groups which will each appear as columns in the statistics tables. The bottom of the “Edit Labels” tab is used to set up labels within the selected group. To assign a label to an object one has to select the object in the 3D View and then click the “Assign” button for the respective label.
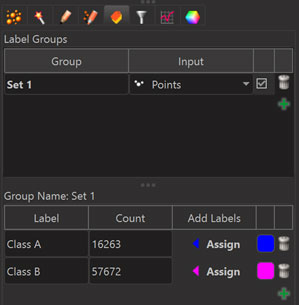
Note that each object can receive only one of the different labels within a label group. To assign multiple labels to one object it is necessary to create multiple groups.
In Imaris 9.6 the Vantage 1D View lets you split values by their labels to facilitate a simple visual comparison. This makes it very simple to see statistical differences between classes of objects. For example if you have classified Spots as being inside or outside of Surfaces, you can easily compare any statistics of these two groups in a side-by-side plot in Vantage1D.
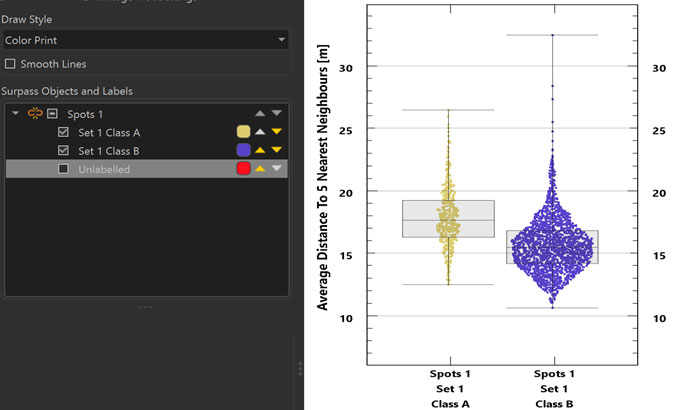
The “split” of a plot into several plots for different labels (if objects have been assigned different labels) is accomplished by clicking on the “chain” icon in the “Surpass Objects and Labels” user interface.The “triangles” in the “Surpass Objects and Labels” user interface facilitate changing the sequence of the plots.
Imaris 9.6 has a new module for 2D scatterplots. The 2D scatterplots let you quickly display statistical relationships between two variables. The following 2D scatterplot for example shows a clear relationship between “Speed” and “Position X Reference Frame”.

To further analyze such relationships you still have to export statistics from Imaris and work in a statistics software but within Imaris Vantage 2D you can quickly visually inspect your data before you start a statistical analysis in another software.
Imaris 9.6 has a new color picker that pops up when you click on a color button.

Clicking on the triangle next to the Color bar enlarges it to provide access to more details.

The 3DView of Imaris 9.6 allows navigation and selection of objects without the mode-switch (Esc-Key) that was necessary in previous imaris versions. In the new version Mouse-Press+ Drag generally leads to navigation, unless the mouse click hits a selected object with a manipulator such as a clipping plane or a reference frame. If you do not drag the mouse Mouse Release is the action that leads to component selection and object selection. To manipulate Ortho Slicers, Clipping Planes, and other objects with manipulators these objects must be selected prior the Mouse-Press + Drag for manipulation.
In some rare situations the new behaviour may be challenging. In those cases you can use the Alt-key to force navigation.
Imaris 9.6 clearly distinguishes Surpass component selection in 3D View from selection of objects within a Spots/Surfaces/Filaments/Cells component. When you left-click on a component that is not currently selected, then the only thing that happens is that the component will become the selected component. Nothing else. When you left-click on an already selected Spots/Surfaces/Filaments/Cells component, then the object selection within the component is changed. One could say briefly: “first click selects the component, second click works within the component”. The same goes for other objects such as reference frames. The first click selects them. The second click lets you modify them.
For the purpose of efficient selection of objects within a component we have added a Circle-Select mouse mode next to the “normal” Select mouse mode in the 3D View. In Circle-Select mode the mouse pointer has a circle drawn around it and the entire region of the circle acts as the region for selection of objects. This lets you select many objects with a single click. Using Ctrl+Click you can add all the objects “behind” the circle to the selected objects or, if all objects within the circle are already selected you can remove them from the selection with a Ctrl+Click.
Note that in Circle-Select mode the center of the circle still keeps acting “normally” in the sense that it changes the selected component in the surpass tree when it hits a component that is not currently selected.
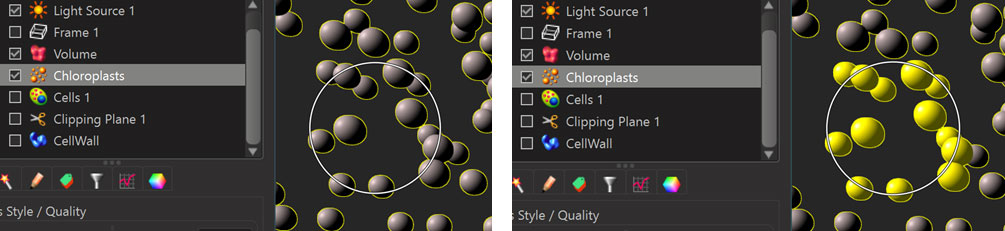
The Esc-key as well as the keys 1, and 2 let you quickly switch between “Pointer Selection Mode”, and “Circle Selection Mode”.
When working with time data and tracked objects in Imaris it can sometimes be useful to select “entire tracks” and at other times it is useful to select individual objects within a track. For example spots may be filtered using the statistics value “Distance to Nearest Neighbour” either with the aim of selecting all spots that have a certain “Distance to Nearest Neighbour” or with the aim of selecting all tracks that contain some spots that have a certain “Distance to Nearest Neighbour”. The former happens when Selection Expansion is set to “Object Selection”. The latter happens when Selection Expansion is set to “Full Track Selection”. The toolbar in the 3DView gives you easy access to the selection expansion choices when you work with time series images.
Imaris 9.6 can open BDV files without conversion to IMS.
Imaris 9.6 supports writing and reading of IMS files with several different compression methods a) Compatible b) Smaller Files, and c) Faster Save.
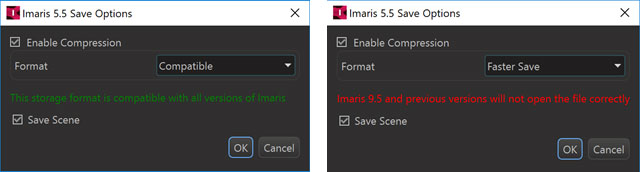
The “Compatible” option uses gzip compression.
The “Smaller Files” option uses HDF5 shuffling in combination with gzip compression.
The “Faster Save” option uses LZ4 compression instead of gzip compression.
Please note that the “Smaller Files” and the “Faster Save” option produce files that cannot be opened with Imaris versions prior to 9.6.
On high resolution displays recent operating systems provide “scaling” functionality that allows users to choose the size of text and button appearance. Imaris 9.6 is the first Imaris version that supports this feature. You can now comfortably run Imaris on high resolution displays with buttons and text in a size that suits you.
Note that the Imaris’ graphics quality and performance are both impacted by high resolution displays. Imaris can render more detail on high resolution displays and this comes at a computational cost. If you change to a high resolution display you should take this into account and attach it to a GPU that is capable of rendering fast enough to that display.
| 64 Bugs Fixed in Imaris 9.6 | |
| 6914 | Changes in the settings tab leads to crash using customer dataset |
| 7650 | Number of time points differs between mac and windows for the same file |
| 9012 | Animation toolbar poorly drawn using "Larger" display scaling in Windows 4K HIGH DPI |
| 10304 | Huge 2D ND2 |
| 10418 | Checking/Unchecking objects doesn't work as expected in Vantage |
| 10743 | New surface makes a rough masked channel |
| 10862 | Annotation check box doesn't work properly |
| 10864 | Selection color lost after toggling display off/on |
| 11328 | Imaris File Converter crashes when converting new multi |
| 11354 | 'ND2 file conversion gives voxel size with rounding error |
| 11367 | Data Volume does not update after changing voxel size |
| 11569 | The manipulator is far away from the image and can't be moved in Cell slicer display mode |
| 11684 | OME XML/TIFF from an Aurox system are 'empty' in Imaris (Bioformats is fine)) |
| 11687 | ImarisCell Slicerviewer is broken when using any ROI in the creation wizard |
| 11745 | Saving image as OME TIFF in Imaris fails for multi |
| 11758 | Imaris cannot correctly read time interval of .vsi files |
| 11761 | Imaris cell fails to recognize empty space in 2D volume |
| 11764 | Impossible to save Coloc channel using "Save" button |
| 11768 | Converted ND2 file is twice as large when compared to 9.2 conversion |
| 11862 | Spot Placement (Surface of Object) fails if a surface was made manually drawn contour lines |
| 11871 | Pressing "Ctrl+Shift+G" closes (crashes?) Imaris without any message |
| 11880 | Correct Drift throws 'out of memory' error for a small timelapse image |
| 11903 | Surface Creation Parameters are shown after Spots Store Parameters for Batch |
| 11925 | Surfaces Preview mode |
| 11956 | Surface preview is very different from actual detected surfaces when the Threshold value is higher than default |
| 11960 | Surface Selection does not work in Batch Setup Step 2 |
| 11967 | Cannot run Imaris 9.5.0 on MacOS Catalina 10.15.0 |
| 11975 | Batch parameters cannot be saved with " / " character in name |
| 11978 | Convert tiff series to ims with Configure File Series does not work when dialog is opened twice before conversion |
| 11980 | Surface center points are all white under Statistics Coded and Object ID |
| 11981 | Shortest distance calculation is triggered whenever Preferences are opened |
| 11983 | Imaris 9.5.0 claims no valid license even if there is one and starts normal |
| 11990 | Right Click in Filament Starting Point Threshold also sets lower threshold |
| 11993 | Material Label Checkbox is present in too many places |
| 12003 | Cells transparency level goes back to default when change Object Type to Nuclei |
| 12005 | Cannot read oib file |
| 12019 | CZI file with multiple tiles converts to ims with only one tile |
| 12022 | Image data saved in Big endian layout in IMS |
| 12028 | Stitcher Demo licenses cannot be activated |
| 12033 | Clicking Batch on non ims file shows error: "Unknown File Format" |
| 12037 | New Color Tab with Label Checkbox is not reflected in reference manual |
| 12040 | File Converter only convert the first tile from .czi datasets |
| 12046 | Annotations remain after opening a different image |
| 12071 | 2D surface display is strange |
| 12072 | Conversion of ICS/IDS files in Imaris9.5 is broken |
| 12094 | ICS(2) import wrong on PC (channels are shifted, Intensities wrong); Mac import is correct |
| 12101 | This tiff file does not convert properly |
| 12102 | ImarisConvert ignores the |
| 12103 | Create multiple contour Surfaces leads to crash |
| 12106 | LIF file conversion in Arena (and FileConverter) fails on PC; Mac works |
| 12107 | Average statistics |
| 12113 | Imaris crash on "Edit" batch (deconvolution) parameter |
| 12123 | Filament data with spines is not able to export average statistics |
| 12125 | ims files with corrupted Surfaces |
| 12142 | Stitcher Installer did not install required Java version |
| 12170 | Scrolling within Image Proc channel tab crashes Imaris |
| 12184 | Crash during batch creation when stepping back from "2 |
| 12197 | MCR 2014b installer does not run on MacOS 10.15.x |
| 12208 | Statusbar shows white border |
| 12224 | Problems with HostID on MacBooks |
| 12238 | Changing Tracks color does not work |
| 12260 | Switching surface visualization from surface to slice view takes too long |
| 12277 | ImarisStitcher cannot stitch in macOS 10.15 Catalina due to Java |
| 12283 | Custom Python 3.7 XTension failing to launch from Imaris 9.5 from Surpass Dialog menu (Gear tab) |
| Fixed 5 Bugs (since Imaris 9.6.0) | |
| 12372 | Crash when going back and forth between step Threshold and Filter Seed Points |
| 12365 | XTensions do not start. Fatal error loading library ... mclbase.dll |
| 12406 | Select Coloc tab with 150% or higher windows scaling crashes Imaris |
| 12511 | Imaris doesn't recognize .ims file converted from .tif since 9.6.0 |
| 12545 | Setting the Date/Time and Format in Windows to anything else than English will cause issues in Wizards entering numbers with decimal point |