Neurons
- Soma Volume
- Branch Points
- Branch Angles
- Spine Density
- Sholl Intersections
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
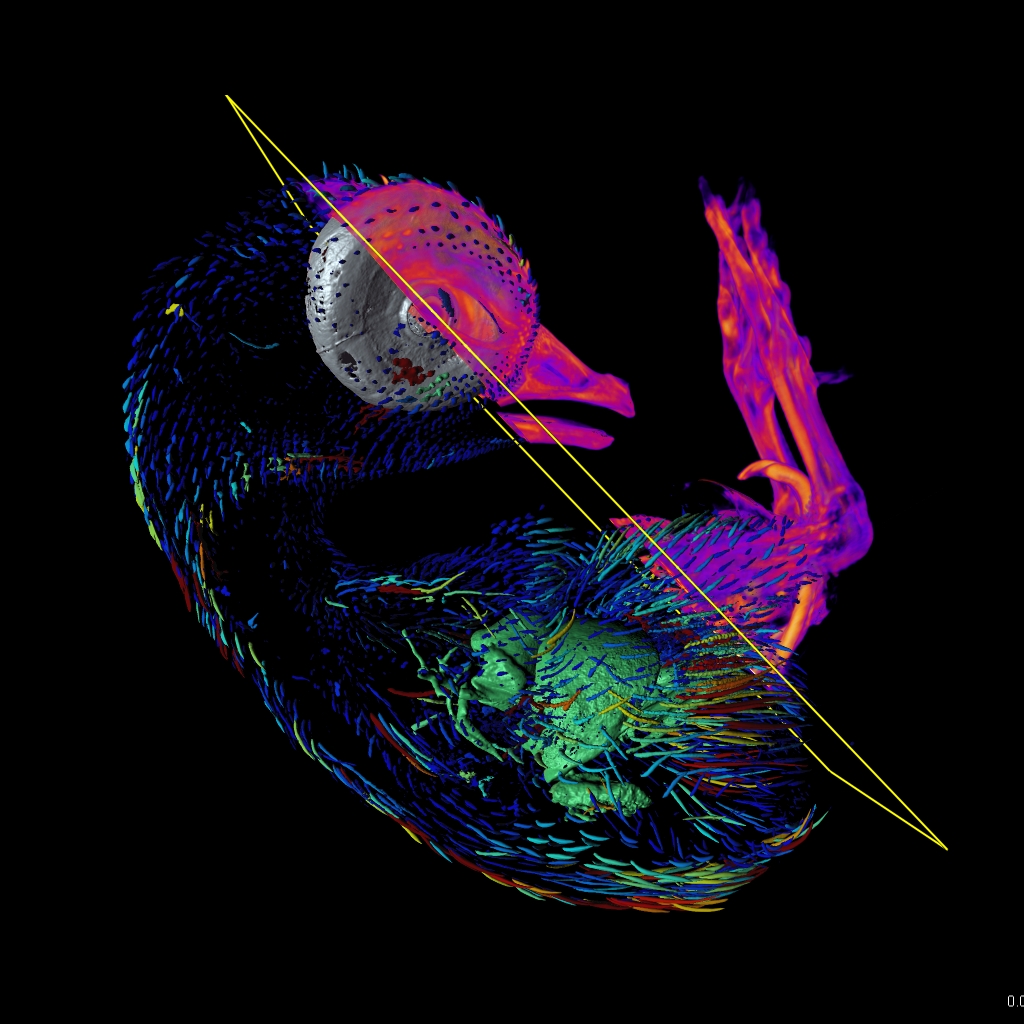
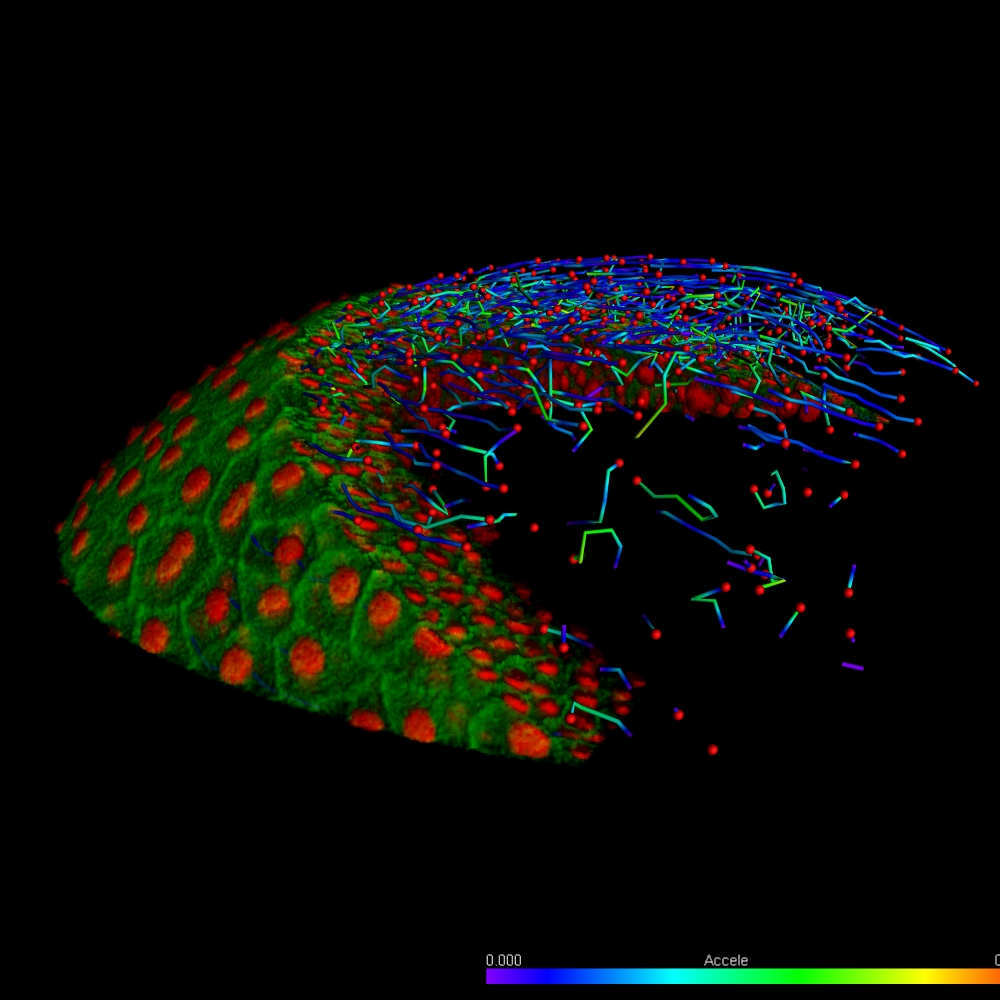
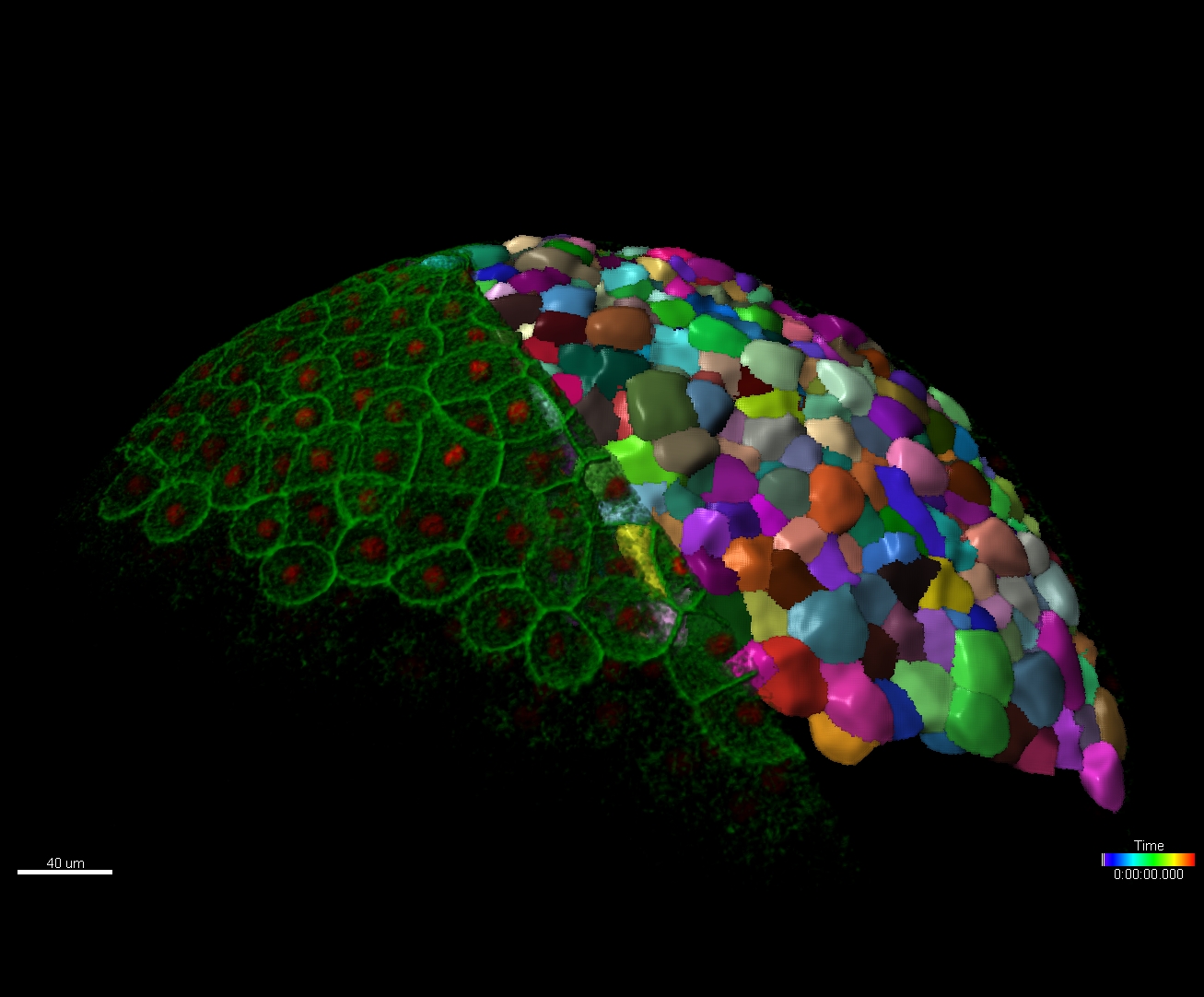
The Imaris for Neuroscientists package enables 3D reconstruction of neurons and arborization analysis. It can resolve various structures, such as axons and dendrites, somas, dendritic spines, microglia, or astrocytes. It calculates a range of neuron-specific measurements, such as dendrite or segment length, orientation, diameter, branch level, spine density and spine shape analysis and classification.
In addition, Imaris for Neuroscientists is perfect for tracing blood vessels and getting measurements like segment length and total length, diameter, mean segment intensity and orientation of segments. In addition, Imaris for Neuroscientists is great for object tracking, plotting, group comparisons and provides a two-way interface for customization in Matlab, Java, or Python.
Request Pricing/Free Trial Existing Customer - Satellite LicenseMicroscopy image analysis requires precise detection of multiple biological structures based on the initial signal saved as an image data. Object detection in Imaris will be quick and easy as users are guided within a wizard driven interface to create perfect models of their data. With our patented Torch™ tool and several performance improvements tracing neurons with Autopath or Autodepth in dense and thick samples is very efficient and becomes a unique experience.
Depending on their function and origin in the brain, neurons vary in shape and size. In addition, changes in neuron morphology can be indicative of various diseases or pathological conditions. The neuronal structure can be as well modified during learning or regeneration. Analysis of the neuronal branches and their structure is a valid neuroscience research tool and can be accelerated with a dedicated 3D image analysis software. Imaris for Neuroscientists offers:
Dendritic spines are small neuronal protrusions, which act as sites of a synaptic contact. Spine distribution (density) and morphology is changing during brain development and throughout the live and is associated to learning ability as well as some neuropsychiatric disorders. Modern deep fluorescent microscopy techniques, such as Dragonfly 600, enable visualizing spines in sufficient resolution and in near physiological conditions, including living neurons. Imaris for Neuroscientists is a perfect image analysis tool for:
Glia cell structure and shape is related to their function and ability to interact with neuronal cells. Dysfunctions of glial cells can be associated with several neurodegenerative diseases. Neuroscientists study glia inside the brain tissue sections from both healthy animals and from transgenic mouse disease models. Due to the complexity of the neuronal glia images, 3D image analysis software is essential for acquiring the quantitative information and comparisons between samples. Imaris for Neuroscientists offers:
Alterations in the brain vascular structure and network is often observed in progress of several cerebrovascular diseases, as well as during the brain regeneration after cerebrovascular events such as stroke. In addition, it’s been reported that the brain vasculature may play an important role in neurodegenerative disorders. Imaris for Neuroscientists enables segmenting and analysis of the vasculature network in the fluorescently labelled tissue sections thanks to:
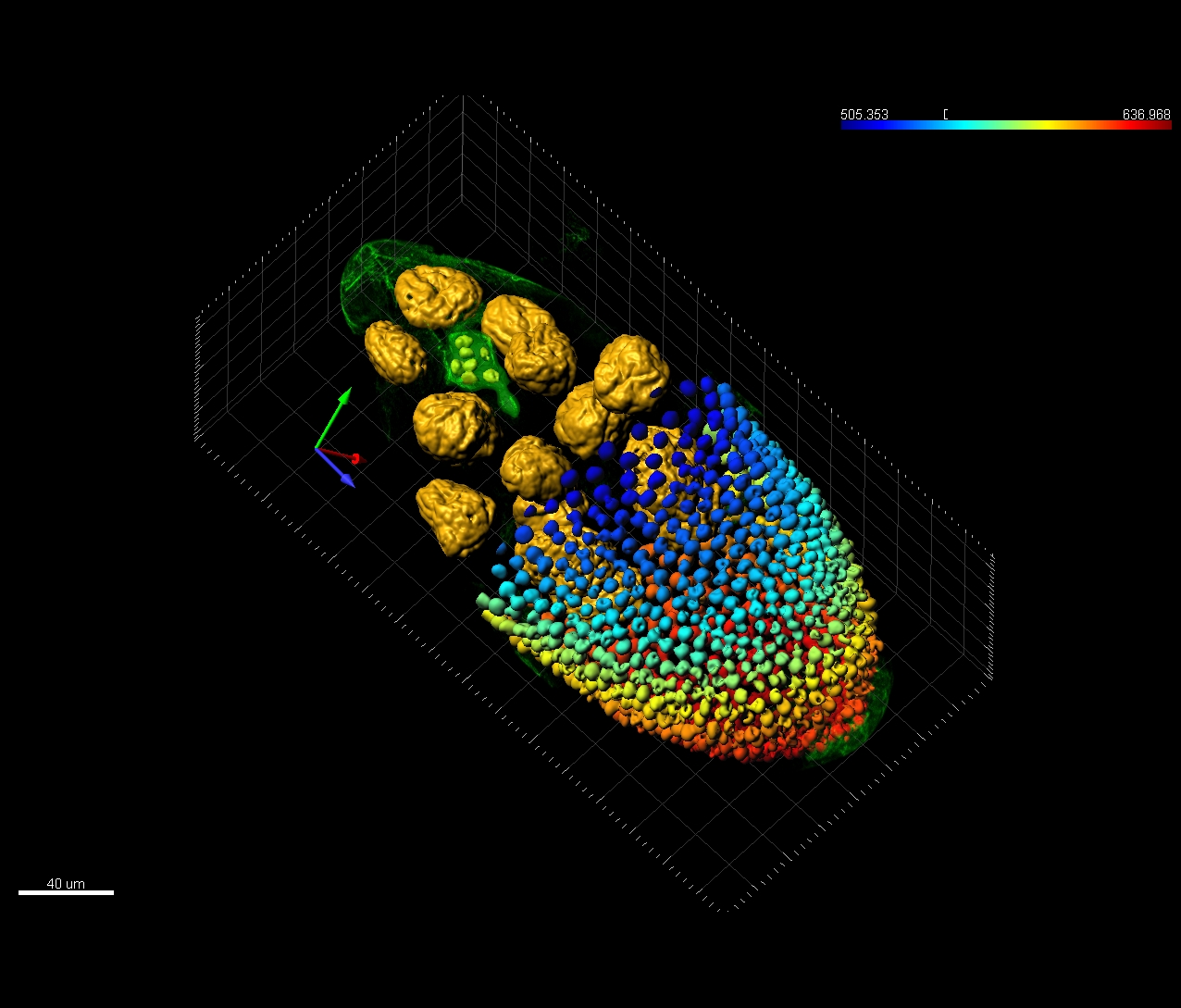
Amyloid plaques accumulation is believed to play a role in the pathology of the Alzheimer disease, yet the mechanism is still unknown. Visualization of amyloid plaques in brain tissue slices or whole cleared brain section is an important research tool and multichannel fluorescent confocal microscopy is a perfect tool for those research, yet requires an advanced image analysis software to quantitatively assess the extent of plaques formation and their distribution in relation to neurons and microglia. Imaris for Neuroscientists can:
Synapse formation and synaptic plasticity is an important part of learning process, while an impaired synapse function can be a feature of some neurological diseases. Quantitation of synapse number and distribution can reveal some important information about the brain circuits and their adaptive or pathologic alterations. Imaris for Neuroscientists can:
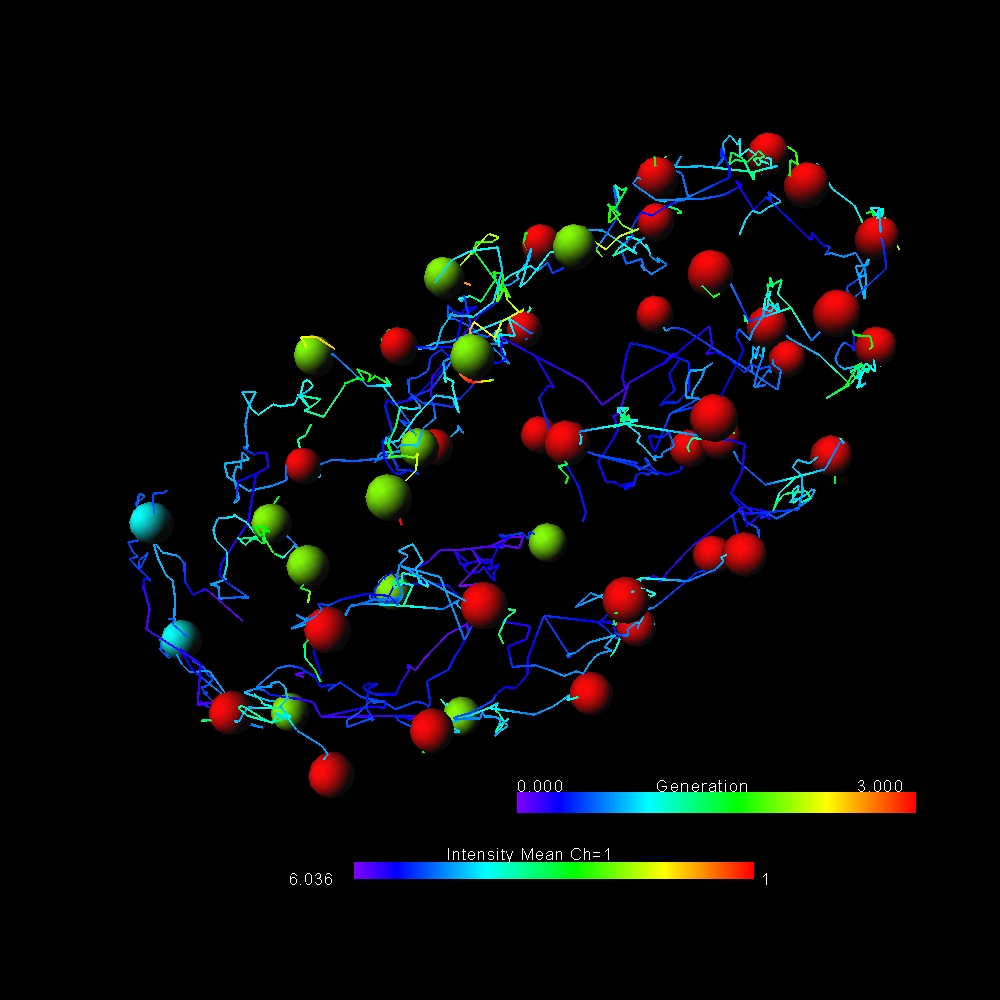
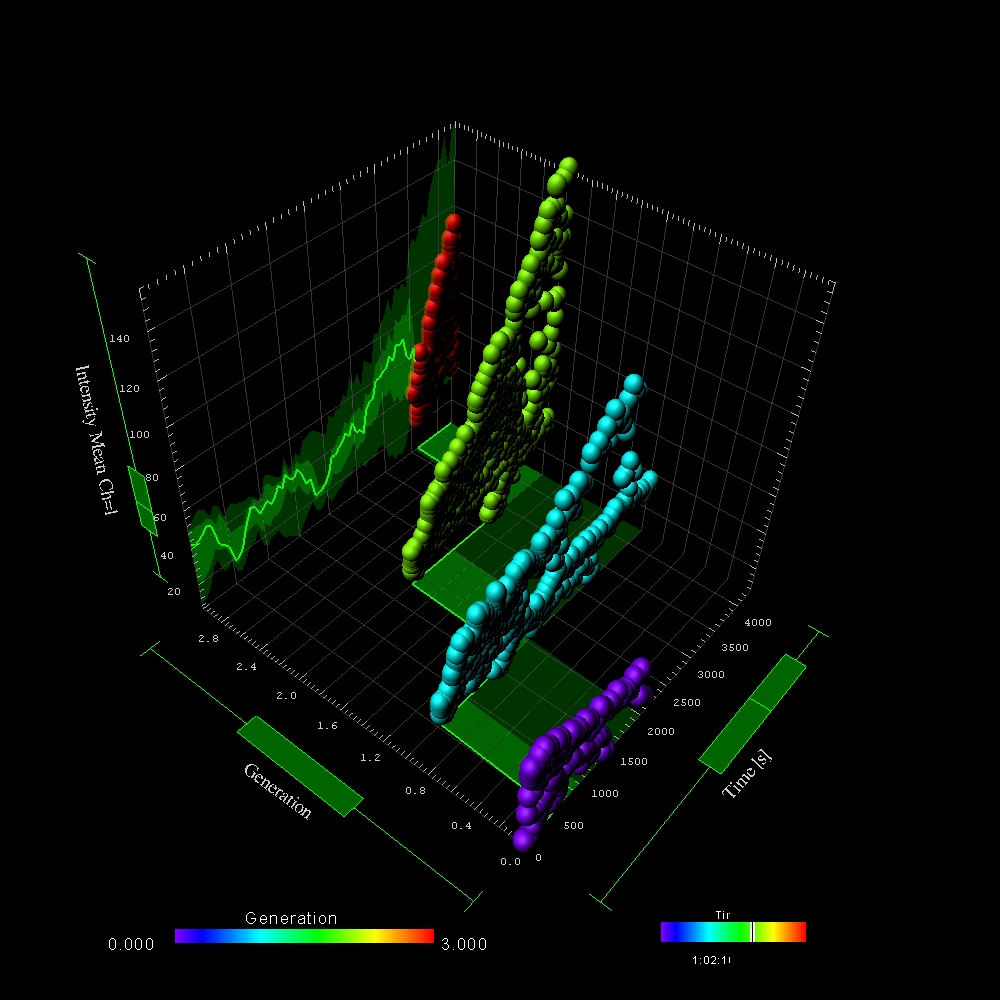
Imaris calculates a wide range of statistics for all detected objects: Filaments, Surfaces and Spots. Users can find a range of neuro-specific parameters, distance and colocalization statistics and morphological measurements. All values can be used for classification, color coding, plotted inside Imaris (using Vantage plots) or exported in an .csv or .xls file format. Parameters presented below are the most common statistic types needed by biologists. Imaris reports many more.
This package is about much more than just tracing. With the inclusion of various Imaris modules, you can experience complete analysis freedom during your experiments.
Automatic neuron tracing combines the benefits of an intensity-based method with a unique machine learning approach, providing the most flexible tool on the market for tracing a multitude of neuron types. Creating filament models is very easy, even for non-experienced users, because they are guided via precisely designed wizards with the result previewed after each step. By following the wizard instructions accompanied with a few paintbrushes along the structures of interest, they are separated from the background. Tracing neurons inside Imaris for Neuroscientists is:
Imaris for Neuroscientists includes a dedicated new Loop algorithm for tracing the blood vessels and get the for the efficient detection of dense vascular networks inside the brain or other organs. The characteristic feature of the blood vessel network is the large diameter difference between the main thick vessels and the small capillaries, so the Imaris 10 for Neuroscientists introduced a new multi-scale seed point feature for accurate detection and measurements. Tracing blood vessels is very easy and efficient, thanks to the combination of an intensity method with the machine learning approach. The benefits of tracing blood vessels inside Imaris for Neuroscientists are:
Semi-automatic tracing tools in Imaris for Neuroscientists are perfect for editing automatically detected neurons or vessels or tracing filamentous structures in big datasets of multiple GBs in size (up to 1 TB tested) and dense networks like cleared samples, cleared tissues or entire brains. User only selects the start and end points in 3D while the best fitting filament path is calculated and highlighted. Tracing can be paused after every intermediate step and started again from the selected end or branching point. The filament diameter is automatically calculated from the structure. Semi-automatic methods in Imaris for Neuroscientists are unique, because:
Regardless of which filament tracing method is chosen (wizard driven automatic tracing or an Autopath/Autodepth mode) dendritic spines can be automatically detected and precisely modelled based on the threshold. The benefits of the automated dendritic spine detection method in Imaris are:
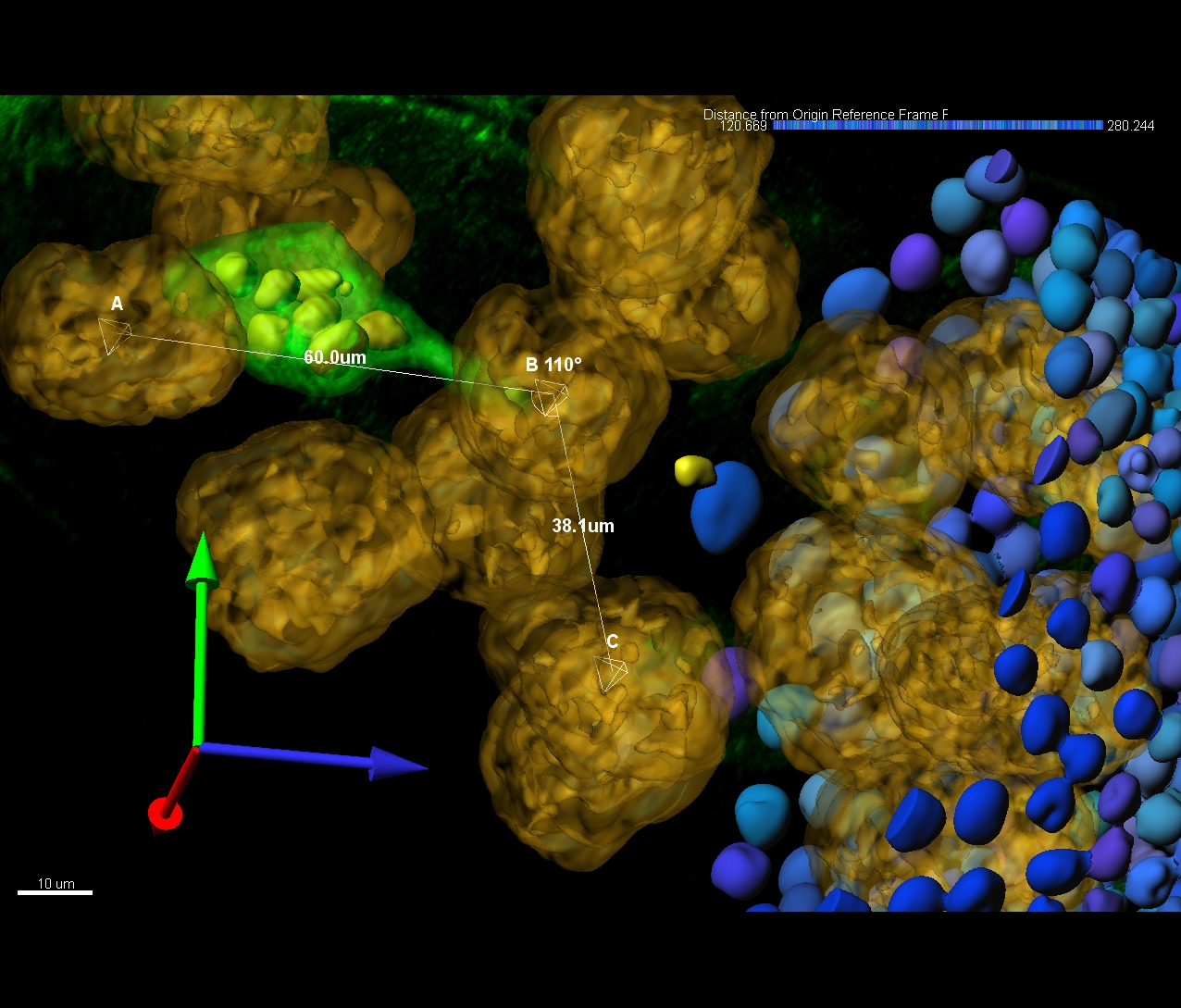
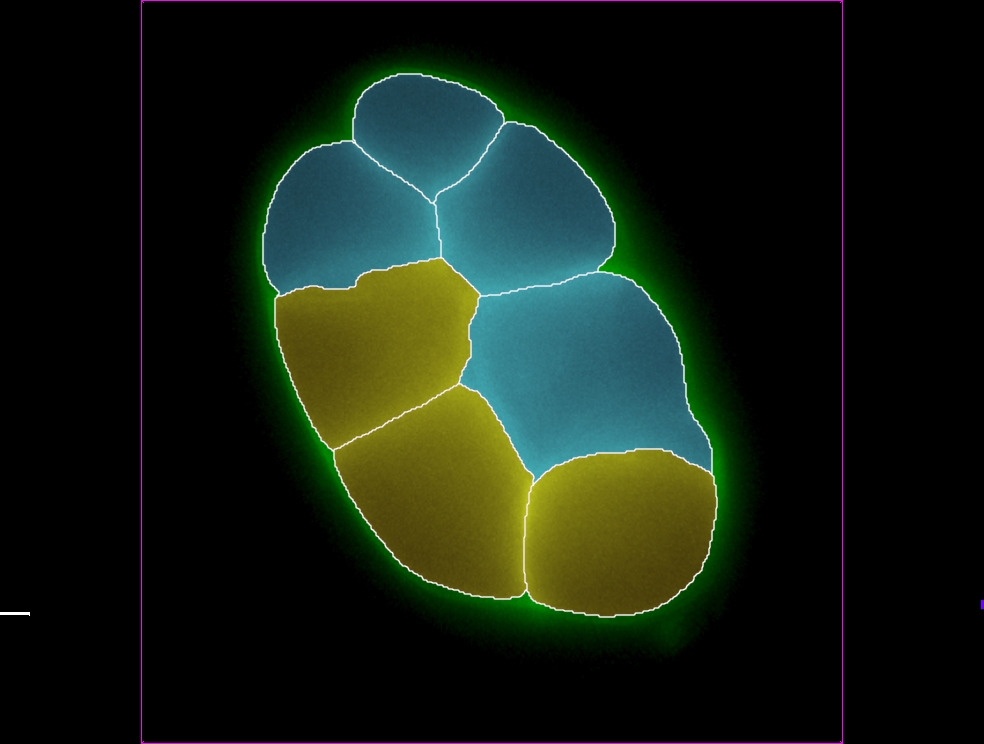
Surface objects can be created over the boundaries of objects of different shapes and sizes, like cells, nuclei, nucleoli, brain structure or a filament while the Spots objects are used to model point-like or vesicle-like structures in the data and to quickly identify thousands of structures. Surface and Spots model are complimentary to neuron tracing and provide additional set of measurements, such as volume, precise position, or intensity. What makes Imaris for Neuroscientists unique is the enablement of automated distance measurements between all the detected structures, including filamentous structures and other stained objects. In Imaris for Neuroscientists you can:
The Imaris Learning Center hosts a wide range of tutorial videos, how-to articles and webinars to guide you through the many features of Imaris. We have provided some links below which will get you started on some of our most recent developments.